Demographics and recruitment
Table of Contents
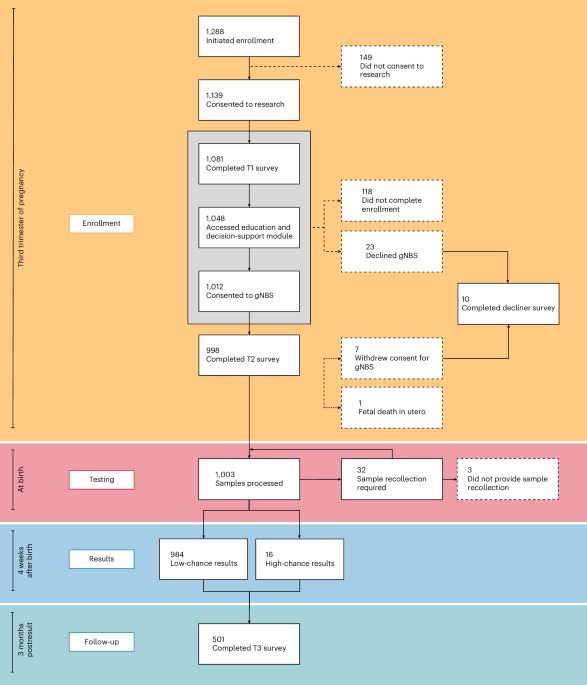
Study enrollment was initiated12 by 1,288 prospective parents, with 301 (23%) not completing enrollment, declining screening or withdrawing from the study. gNBS was completed for 1,000 newborns over 16 months, of whom 523 (52%) were male and 477 (48%) female, including 13 sets of twins (Fig. 1). The participant sample was underrepresented for parents under the age of 30, with two or more children, from regional areas and in Quintiles 1 and 2 of the Index of Relative Socioeconomic Advantage and Disadvantage. A relative overrepresentation of higher-educated individuals was observed (Fig. 2).
Eligible participants initiated enrollment via the online platform Genetics Adviser, accessible via a QR code either on a study invitation card obtained from a healthcare professional (active recruitment) or on a range of other advertising material (passive recruitment). Engagement was tracked from initial login to the platform using a unique study ID. Completion of the Genetics Adviser education and decision-support module, T1 and T2 research surveys, and consent to both research participation and clinical gNBS was mandatory to complete enrollment. Completion of the T3 survey, distributed 3 months postresult, was optional. Solid line, pathway to study completion; dashed line, pathway for participants who were lost to follow-up, declined gNBS or withdrew from the study.
a,bComparison between the study cohort and the Victorian perinatal population in 2022 in relation to age (a) and parity (b), sourced from the Australian Institute of Health and Welfare’s National Perinatal Data Collection. c–fComparisons between the BabyScreen+ cohort and the age- and sex-matched population of Victoria, Australia, based on Australian Census data in relation to geographic location (c), highest level of educational attainment (d), area of ancestral origin (e) and Index of Relative Socioeconomic Advantage and Disadvantage (f). Geographic location (Remoteness Area) (c) and Index of Relative Socioeconomic Advantage and Disadvantage (f) are based on the classifications used by the Australian Bureau of Statistics. These variables are from a level 2 statistical area (average population of 10,000 people). Participants were able to select more than one response when reporting their ancestry. Ancestry data (e) are presented as a proportion of the total responses. The top five ancestries from the BabyScreen+ data are presented with the addition of Aboriginal and Torres Strait Islander ancestry. Percentages are rounded to the nearest whole number and totals may not equal 100. The demographic variables were compared between BabyScreen+ participants whose newborns had genomic newborn screening (n = 987) and the relevant Victorian population datasets using chi-square tests. Significant differences were observed for age (x2 = 150.80, P < 0.001), parity (x2 = 69.04, P < 0.001), geographic location (x2 = 72.46, P < 0.001), highest level of educational attainment (x2 = 901.36, P < 0.001) and Index of Relative Socioeconomic Advantage and Disadvantage (x2 = 37.69, P < 0.001). aThe BabyScreen+ demographic survey indicated that Oceanic ancestry refers to people from the Pacific Islands or Micronesia. The 2021 Australian Census did not provide such guidance and as such it is likely that many of the responses indicating Oceanic ancestry refer to respondents of European ancestry born in Australia. Due to the discrepancy in how this variable is ascertained, a chi-square test was not run for this variable.
Recruitment by a healthcare professional (active recruitment) resulted in the highest number of study participants completing enrollment (667 out of 998, 67%, Fig. 3), with social media advertising the most successful passive recruitment method (190 out of 998, 19%). Active recruitment had the lowest proportion of incomplete enrollments (Extended Data Table 1). A detailed breakdown of cohort demographics by recruitment method is provided in Extended Data Table 2 and Supplementary Fig. 1.
Each participating family is represented as a single square, with n = 987 enrolled participants whose newborns had genomic newborn screening including 13 sets of twins. Recruitment methods were broadly categorized as ‘active’, where a study invitation card was offered to a prospective participant by a healthcare professional (midwife, obstetrician or primary care physician) in both the public and private healthcare sector. Passive methods were only utilized in the public healthcare setting and included study posters and videos in antenatal clinic waiting rooms, study invitation cards enclosed in hospital antenatal packs provided by mail, direct text messages to pregnant women, and advertisement on a designated pregnancy app used at one participating hospital. A paid social media advertising campaign on Facebook and Instagram was targeted to women between the ages of 18 and 45 located in the state of Victoria, Australia. The study team recruited eligible participants who contacted the study team directly and unsolicited. Commencement of deployment was asynchronous for different recruitment methods, with major deployment dates indicated.
Feasibility of genome sequencing from dried blood spots
DBS cards from 1,003 newborns were processed for gNBS. Reprocessing was required for 82 (8.2%) samples due to sample-related (3.2%) or process-related (5.0%) failures (Extended Data Table 3). DNA reextraction was performed for 79 (7.9%) samples, using additional punches from the existing DBS card (50), collection of additional DBS cards (7) or collection of fresh blood samples (22 samples, Extended Data Table 4). Three participants declined sample recollection.
Sequencing data generation failed target coverage requirements for 191 (19%) samples, requiring additional sequencing. Process improvements, including optimization of sample quantitation methods and sequencer loading concentrations, reduced failure rates from an initial average of 28% to less than 5% (Supplementary Fig. 2).
All reextracted and reprocessed samples were successfully reported on the second attempt. Excluding samples that required reextraction would have led to two missed high-chance results (UNC13D and GnasTable 1).
Average turnaround time (TAT) for stdNBS followed by gNBS was 24 days from sample collection (target: 28 days, minimum: 11, maximum: 68, 95% confidence interval (CI) 23.5–24.5). Average TAT for gNBS was 13 days (minimum: 6, maximum: 56, 95% CI 12.7–13.3), with 73% reported within the target of 14 days. The proportion of reports issued within target TAT increased from 65% to 81% following reduction in sequencing failure rates.
Genomic data analysis and interpretation
Analysis protocols (Fig. 4) were validated using a cohort of 108 clinical cases of critically ill infants undergoing genomic testing, with 61 known low-chance and 47 known high-chance results2. Validation data showed >97% sensitivity with 46 of the known high-chance cases correctly flagged. The high-chance case missed by the analysis had a multinucleotide variant (ACAD9, NM_014049.4c.1376_1381delinsCCT, p.(Lys459_Ser461delinsThrCys). This was incorrectly annotated by the analysis software as two non-high impact variants, a known limitation. No cases were incorrectly classified as low-chance by the automated system; however, 32 of 61 (51%, Extended Data Table 5) of known low-chance cases required manual review of nonreportable variants.
aKnown benign variants include those classified as benign or likely benign by the Victorian Clinical Genetics Services (VCGS) and/or submitted as such in national (Shariant39) and/or international databases (for example, ClinVar, (likely) benign with at least two stars). bVery common variants include those with a frequency of more than 2% in VCGS internal data and/or the Genome Aggregation Database. cKnown pathogenic variants include those classified as (likely) pathogenic by VCGS, in national databases (Shariant39) and/or international databases (ClinVar). dHigh-impact variants include nonsense, frameshift, canonical splice site, start loss, stop loss, and large deletions and duplications. eVariants in the literature include missense, in-frame and splice region variants in the Mastermind Cited Variant Reference (now known as the Indexed Variant File). fVariants too common for monoallelic disease (0.01% in Genome Aggregation Database) or biallelic disease (0.1% in Genome Aggregation Database). Mode of inheritance (MOI) is derived from the MOI of the gene in the BabyScreen+ panel in PanelApp Australia. gOnly heterozygous variants are allowed for monoallelic MOI. Biallelic MOI requires a homozygous or hemizygous zygosity, or else two (or more) variants in the same gene. VCF, variant call file.
Of the 1,000 study samples, 451 (45%) were automatically reported as low-chance. In the remaining 549 (55%) cases, 1,045 variants were manually reviewed, with 36 variants undergoing full assessment and 18 ultimately reported. Of the nonreportable variants, 1,009 were discarded following rapid manual assessment due to insufficient evidence for pathogenicity, including all copy number variants (Extended Data Table 6). The 18 variants that underwent full assessment but were not reported were excluded for a range of reasons including mismatched mechanism of disease or mode of inheritance, or association with adult-onset or mild disease (Extended Data Tables 6 and 7).
Modeling filter conditions showed an automatic low-chance report rate of 82% was achievable (Extended Data Table 8) but increased the likelihood of false negative results. Only reporting variants previously described as pathogenic in ClinVar would have resulted in the loss of one high-chance result (GnasTable 1).
High-chance results were issued for 16 newborns (1.6%, Table 1). No discordances with stdNBS were identified. One newborn was identified as having hypothyroidism through stdNBS. gNBS provided the precise etiology (Gnas variant) enabling cascade testing and surveillance for multisystem involvement. All high-chance results were confirmed by orthogonal testing on fresh samples, with no discordances. All variants in recessive genes were confirmed in trans.
Clinical impact of high-chance results
Clinical impact ranged from instituting preventative measures or surveillance to active management. Nine results in three genes (G6PD, MT-RNR1 and Ryr1) prompted preventative measures (Table 1). These were managed by the gNBS team without involvement of specialist services. Five results (FBN1, Gnas, ENG, Gb2 and DICER1) led to initiation of surveillance measures, which included echocardiogram, blood tests, MRI scan, audiology assessment and follow-up with specialist physicians. Two results (PKHB and UNC13D) resulted in immediate treatment (Table 1). The infant with glycogen storage disorder due to PKHB variants had unrelated congenital heart disease and had been scheduled for cardiac surgery. Identification of an underlying metabolic disorder enabled appropriate multidisciplinary management of perioperative fasting to avoid episodes of hypoglycemia. The infant diagnosed with UNC13D-related hemophagocytic lymphohistiocytosis (HLH) was clinically well at the time of gNBS result disclosure, but immunological testing revealed early signs of immune dysregulation. The diagnosis enabled early commencement of therapy with immune modulators and proactive planning for bone marrow transplantation, which was performed at 4 months of age.
Testing in first-degree relatives resulted in 20 additional diagnoses (12 parents and 8 siblings). None of the affected relatives were previously suspected of having a genetic condition, although clinical findings and family histories consistent with the diagnoses were present in 4 parents (FBN1, Gnas, ENG and DICER1). Two participants required reanalysis of gNBS data for diagnostic purposes within the study period. One was a newborn with a low-chance result who had a diagnosis of moderate bilateral sensorineural hearing loss from newborn hearing screening. Diagnostic reanalysis did not identify a genetic cause for hearing loss. The second newborn with a high-chance result for G6PD deficiency had diagnostic reanalysis following admission to neonatal intensive care with multiorgan failure. A reanalysis report was issued within 24 h of the request, with no additional genetic diagnosis identified.
Out of 1,012 parents who consented to gNBS, 998 (99%) completed a survey (Fig. 1). We conducted 48 interviews with 46 birth parents and 3 partners. This included 22 pretest gNBS acceptors, 2 gNBS decliners, 17 low-chance results recipients and 8 high-chance results recipients.
Most survey respondents (80%) indicated they consented ‘immediately’ using Genetics Adviser, with the decision perceived as either ‘easy’ or ‘very easy’ (Supplementary Table 1). Only 8% found decision-making difficult. Interviewees valued Genetics Adviser, with education content generally used to reaffirm decisions, provide guidance on what to consider, understand the impact of their decision or facilitate discussions with partners.
Interviewees also described how they considered clinical, psychosocial and practical factors in gNBS decisions. They weighed up benefits of screening, the types of conditions being screened, potential barriers and their ability to navigate results.
Decisions to have gNBS were most strongly influenced by a desire “To know what to expect for my baby’s future” (77% survey respondents) (Fig. 5). The main influence for declining gNBS (10 surveys completed) was concern about the result having negative impact on parents (80%) (Supplementary Table 2).
Older age (for ages 30–34, odds ratio (OR) 2.41, 95% CI 1.35–4.33; ages 35 and over, OR 2.58, 95% CI 1.38–4.81), English as the main language spoken at home (OR 1.9, 95% CI 1.02–3.54) and prior experience with a genetic test (OR 1.8, 95% CI 1.09–3.10) were factors that were associated with participants choosing to have gNBS (Extended Data Table 9). At enrollment, the median trait anxiety score was 32.63 (interquartile range 28.42–38.94). Most survey respondents scored under the cutoff for probable clinical state anxiety at enrollment (80%) and at the T3 survey (85%, n = 501). Postresult return, decision regret was very low (median 0, interquartile range 0–10, n = 506). Parents of infants who received a high-chance result were asked to complete an adapted version of the Genomics Outcomes Scale (GOS). Eight participants (50%) responded to the GOS scale, with a mean empowerment score of 26.3 (95% CI 23.98–28.52) out of 30. Interviewees who received a low-chance gNBS result reported positive impacts such as reassurance. Interviewees who received a high-chance result valued results due to their clinical utility. Prompt genetic counseling and access to high-quality information facilitated adaptation.
Most respondents (80%) would choose to have gNBS for a future baby, and 92% would recommend it to a family member (Supplementary Table 3). All but one respondent thought that gNBS should be available to all parents, and 97% thought it should be publicly funded.