optimizing Automated Streaking Patterns for Single-Colony Isolation in Microbiology
Table of Contents
By Dr. Alice Roberts | 🔶 DATELINE_LOCATION – 2025/08/18 08:04:46
Single-colony isolation is fundamental for microbiological diagnostics, impacting species identification and antimicrobial susceptibility testing. A previous study has reported that manual streaking methods exhibit variability owing to differences in individual techniques, which can compromise the reproducibility of bacterial isolation7. While direct assessments of inter-technician variability in microbiological testing are limited, comparisons between manual and automated streaking systems suggest that automated systems improve reproducibility, enhance single-colony isolation, and increase pathogen detection compared with manual methods, reinforcing the role of automation in standardizing microbiological workflows7,8,11. however, few studies have systematically evaluated multiple pre-installed streaking patterns within a single automated system to determine their impact on single-colony isolation11.
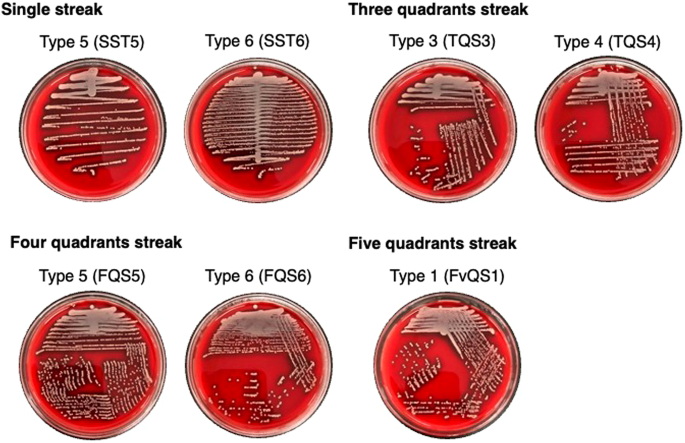
First, we evaluated the performance of seven preinstalled streak patterns in an automated streaking system, focusing on their ability to achieve single-colony isolation from urine cultures across varying bacterial loads. Sst6 consistently yielded the highest number of single colonies among the seven streak patterns tested using standard strains. We selected two single streak patterns (Sst5 and Sst6) for the standard strains and observed important differences in their single-colony isolation efficiencies. Sst5 yielded substantially fewer single colonies than Sst6 for S. aureus, E. Faecalis and E. coli (Fig. 2a) and overall (Fig. 2b). Fqs5 ranked second in the single-colony isolation and exhibited a significantly higher number of single colonies than the four-quadrant streak pattern Fqs6 (Fig. 2b). A previous study comparing two different streak patterns of the WasP system demonstrated that Sst6, which used 27 zigzags, produced significantly more single colonies than single Streak Type 2, which had only 11 zigzags, highlighting the importance of zigzag frequency in optimizing isolation efficiency11. Consistently, our study demonstrated that for both single-streak and four-quadrant streak patterns, a pattern with more zigzag strokes resulted in a higher number of single colonies (Sst6 > Sst5 and Fqs5 > Fqs6). These findings and our results suggest that the streaking pattern selection, especially the number of zigzags, is a key factor influencing single-colony isolation efficiency, reinforcing the need for careful optimization of automated streaking protocols in microbiological workflows.
Although Sst6 demonstrated a superior single-colony isolation efficiency when tested against standard strains, its performance varied when applied to clinical urine samples. Among the three streak patterns, Sst6 showed the highest number of single colonies at bacterial loads from 104 to 106 cfu/ml). Still, at bacterial loads ≥ 10 cfu/ml, Sst6 exhibited a significantly lower single-colony isolation efficiency than Fqs5 (Fig. 3a). This suggests that, even though Sst6 is optimal under moderate bacterial loads, it may not be suitable for cases of heavy bacterial growth, where excessive colony density can interfere with proper isolation. Additionally, Sst6 failed to isolate single colonies of the five species in the four polymicrobial urine samples (Table 1). This finding highlights a critical limitation of Sst6, namely, its reduced efficiency in separating different bacterial species in the presence of multiple organisms. In contrast, Fqs5 demonstrated a significantly better single-colony isolation performance than Sst6 at high bacterial loads, suggesting that it might potentially be a more appropriate choice for such conditions. Among the three streak patterns, Fvqs1 exhibited the lowest coefficient of variation (Fig. 3b). This suggests that Fvqs1 provides the most consistent single-colony isolation across different bacterial loads,making it an ideal choice for standardized clinical workflows.
These findings indicate that although Sst6 was the most effective streak pattern when tested with standard strains, the results from clinical samples indicate that Fqs5 or Fvqs1 may be more suitable choices. Among the evaluated patterns, only Sst6 is suitable for semi-quantitative urine culture interpretation, which is essential for clinical diagnosis based on cfu thresholds. In contrast, Fqs5 and Fvqs1 are better suited for qualitative assessments, such as categorizing growth as rare, some, or numerous, and are more appropriate for samples like pus or wound swabs, where semi-quantification is not routinely performed. The discrepancies between standard strains and clinical isolates highlight the importance of validating streaking patterns in real-world clinical scenarios to ensure optimal single-colony isolation across diverse bacterial loads and sample complexities. Even though local practices may vary, our findings provide reproducible evidence that can support laboratories in selecting streaking patterns tailored to sample type and expected bacterial load. Such data-driven selection is particularly valuable in environments adopting total laboratory automation and Ai-supported colony recognition.
Several studies have examined the single-colony isolation performance of the Copan wasp system3,4,5,6. However, most of these studies primarily compared manual streaking and automated methods or different automated systems. Two previous studies have compared two single-streak patterns within the Copan WasP system but did not evaluate three-, four-, or five-quadrant streak patterns, nor did they comprehensively assess multiple preinstalled streaking patterns5,11. Another study investigated multiple streaking patterns, but each pattern was applied to different sample conditions, making direct comparisons across streaking patterns difficult6. This is the first study to systematically evaluate multiple preinstalled streak patterns within the Copan WasP system using the same set of clinical samples. Unlike manual streaking, in which technician-dependent factors such as inoculum density, streaking pressure, and angle introduce inconsistencies, automated systems minimize variability and improve reproducibility across laboratories12. This study provides critical insights into the effect of streaking pattern selection on single-colony isolation efficiency and offers practical guidance for laboratories using the Copan WasP system. Automated streaking systems are integral components of Tla that enhance the standardization and efficiency of microbiological workflows. With advancements in Tla, Ai-driven colony recognition and digital plate reading are increasingly integrated into microbiology laboratories, streamlining culture interpretation and result reporting. These systems incorporate image analysis software to automatically interpret colonies on agar, thereby reducing processing time while maintaining diagnostic accuracy13. While Ai improves colony detection and species identification efficiency, its accuracy relies on well-isolated single colonies. Studies have shown that ai-assisted microbiology benefits significantly from optimized streaking patterns because suboptimal streaking can result in overlapping colonies,thereby reducing the accuracy of Ai interpretation14,15. This study underscores the critical role of selecting appropriate streaking patterns in automated systems incorporating Ai, ensuring optimal single-colony isolation, improved diagnostic precision, and enhanced reproducibility across laboratories.
This study had several limitations. First, this study focused exclusively on urine samples, which may not fully represent the variability in other clinical sample types. Different streaking patterns may yield different results for samples with more complex microbial compositions, such as respiratory samples or wound cultures. The use of only urine samples was a deliberate design choice to ensure consistency in bacterial load, specimen handling, and clinical interpretation. Urine cultures are among the most standardized and high-throughput specimen types in clinical microbiology, making them ideally suited for automated systems. By focusing on urine, we were able to reduce inter-sample variability and ensure a fair comparison of streaking patterns under controlled conditions. Second, this study was conducted using a single Copan WasP system at a single institution, which limits the generalizability of the findings across different laboratory environments. While the results are likely to apply to other laboratories using the same system,inter-laboratory validation studies are needed. Third, this study did not evaluate the effect of different agar media on bacterial colony morphology and growth characteristics. Future studies should explore the effects of cultural media on streaking performance.Fourth, the comparison between manual and automated streaking in this study was limited to a small number of polymicrobial urine samples (Table 1). While manual streaking yielded more single colonies in some cases, these results may reflect the skill of experienced technologists, which is difficult to quantify or reproduce. Although manual methods may outperform automation in specific instances, the WasP system provided consistent outcomes, supporting its utility for reproducibility and laboratory standardization.
this study evaluated seven pre-installed streak patterns in the Copan WasP system and demonstrated that the single-colony isolation efficiency varies based on the bacterial load and sample composition. Specifically, Sst6 was the most suitable for semi-quantitative interpretation of urine cultures, essential for clinical diagnosis based on cfu thresholds.In contrast, Fqs5 and Fvqs1 provided superior performance under high bacterial load and polymicrobial conditions and were better suited for qualitative assessments. These findings emphasize the need to select appropriate streaking patterns to enhance the reproducibility of automated microbiology.
Moreover, this is the first systematic comparison of multiple pre-installed streaking patterns using the same standard and clinical samples within a single WasP system. The results reveal that performance varies significantly depending on bacterial load and sample complexity,underscoring the practical importance of evidence-based pattern selection. While this study does not introduce mechanistic novelty,it provides essential data to support standardization efforts in laboratories implementing Tla and ai-assisted plate reading systems.
Explainer: Automated Streaking in Microbiology
Automated streaking systems, like the Copan WasP, are used in microbiology labs to prepare samples for analysis. These systems use different streaking patterns to isolate single colonies of bacteria, which is crucial for accurate identification and testing. This study explores which patterns work best under different conditions.
Key Numbers
- 7: number of pre-installed streak patterns evaluated.
- Sst6: Streak pattern most suitable for semi-quantitative urine culture interpretation.
- Fqs5 & Fvqs1: Streak patterns with superior performance under high bacterial load.
Sources
- Previous study on manual streaking variability7
- Comparisons between manual and automated streaking systems7,8,11
- Studies on Copan WasP system3,4,5,6
- Studies on Ai-assisted microbiology14,15